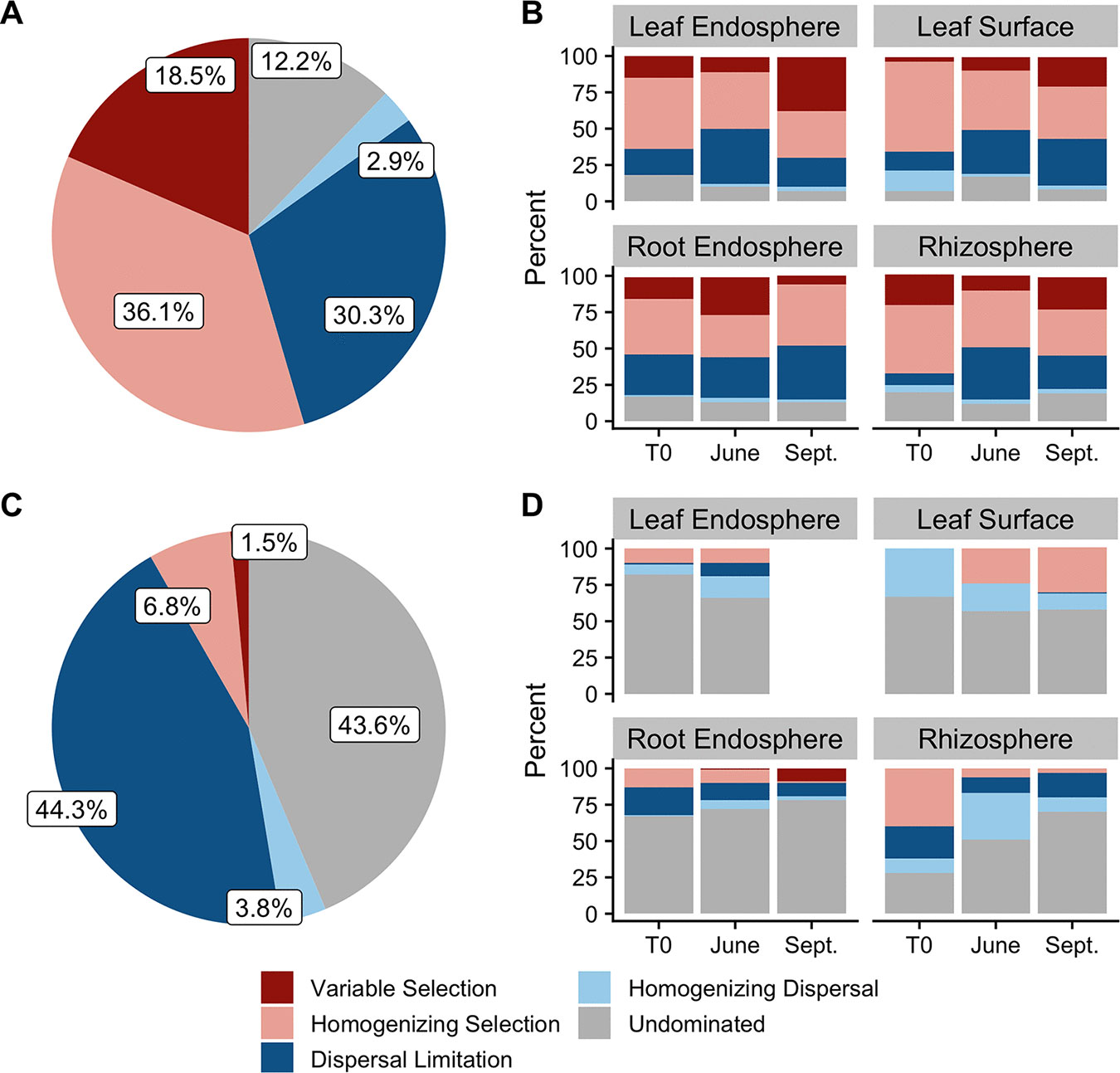
Time, More than Genes, Shapes the Poplar Tree Microbiome
Ecological assembly and source tracking models characterize the initial assembly of the poplar microbiome across plant-associated habitats.

Ecological assembly and source tracking models characterize the initial assembly of the poplar microbiome across plant-associated habitats.
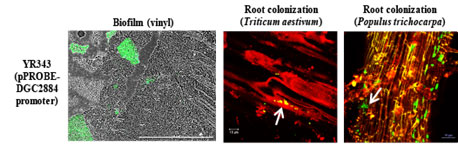
Identification of an enzyme that microbes deploy in the presence of plants leads to discovery of candidate genes involved in root colonization.
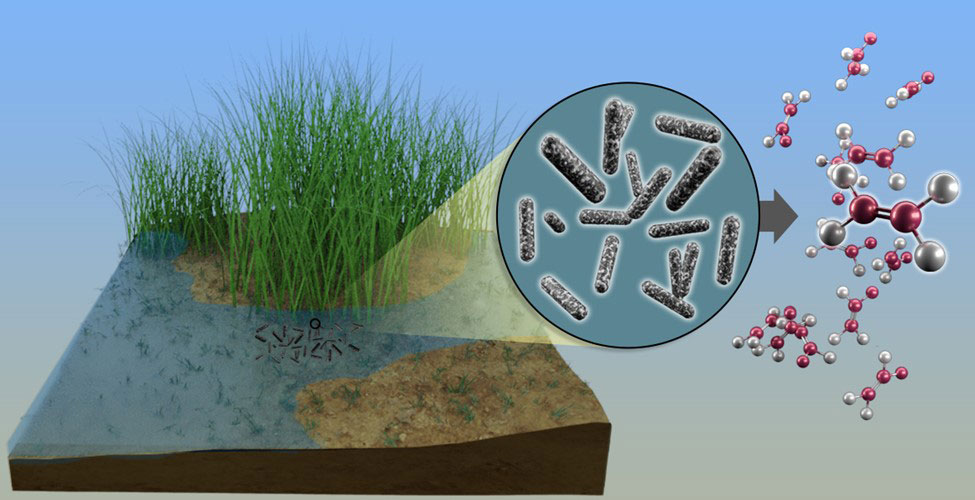

An enzyme system frees sulfur from small organic compounds to make a surprising gaseous side product.
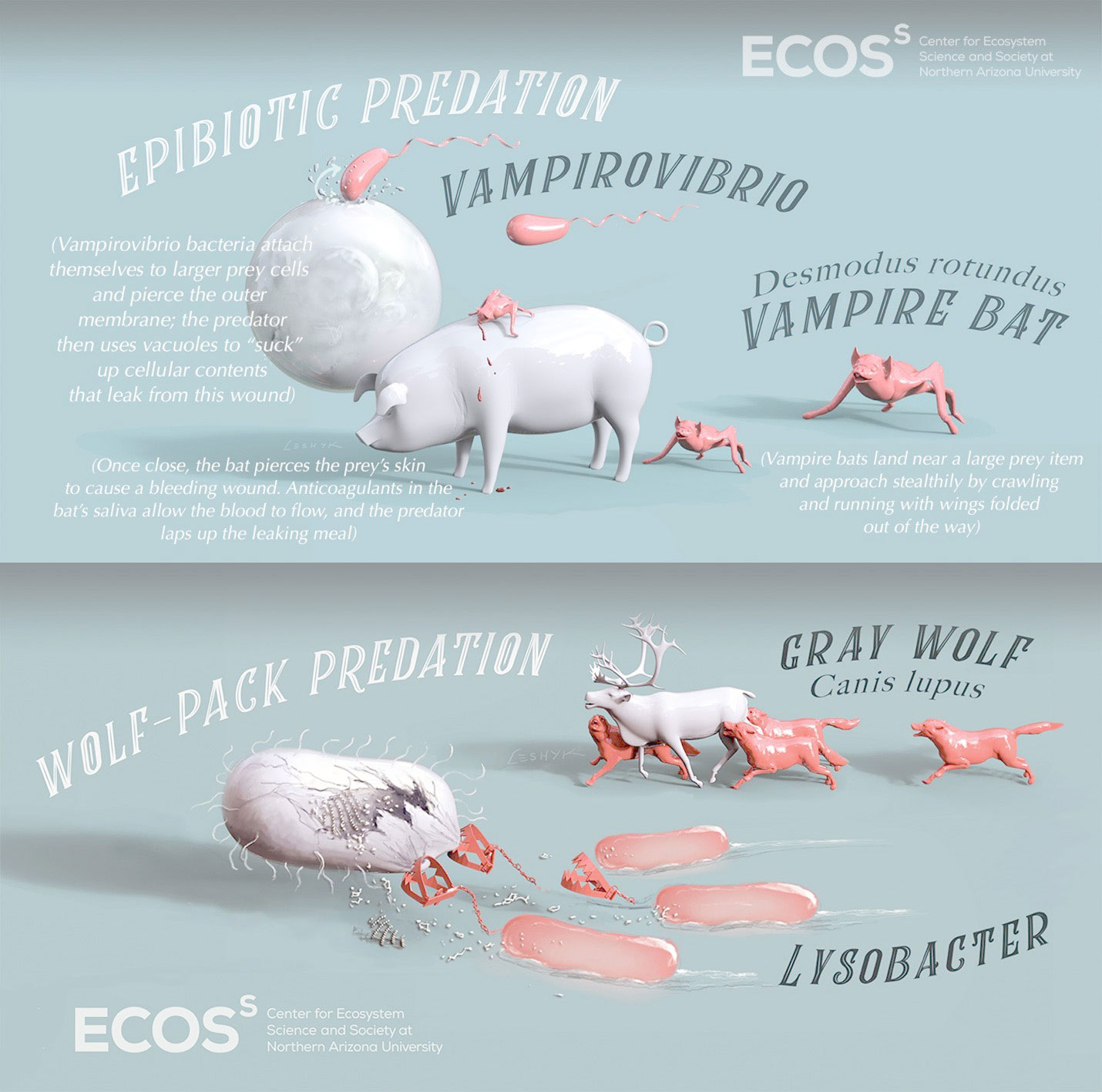

In natural soil, predatory bacteria grow faster than their prey.
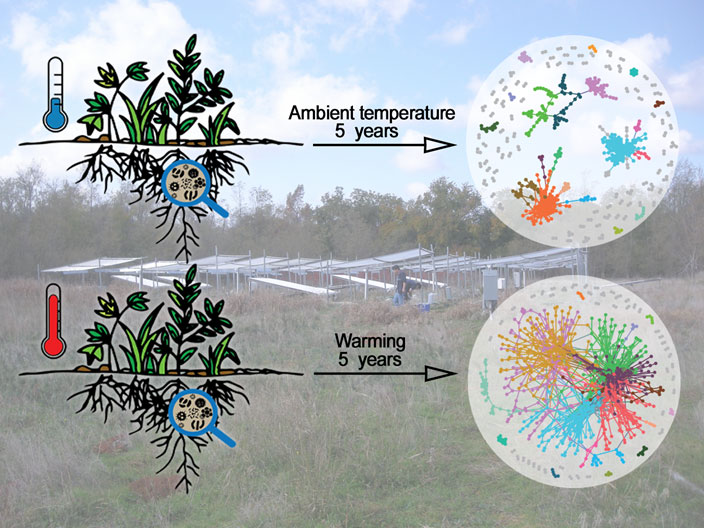
Soil warming leads to more complex, larger, and more connected networks of microbes in those soils

Molybdenum Limits Microbes’ Ability to Remove Harmful Nitrate from Soil
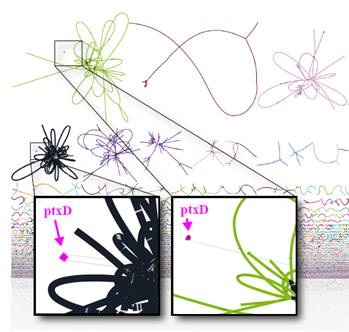

Microbial cycling of phosphorus through reduction-oxidation reactions is older and more widespread than expected.
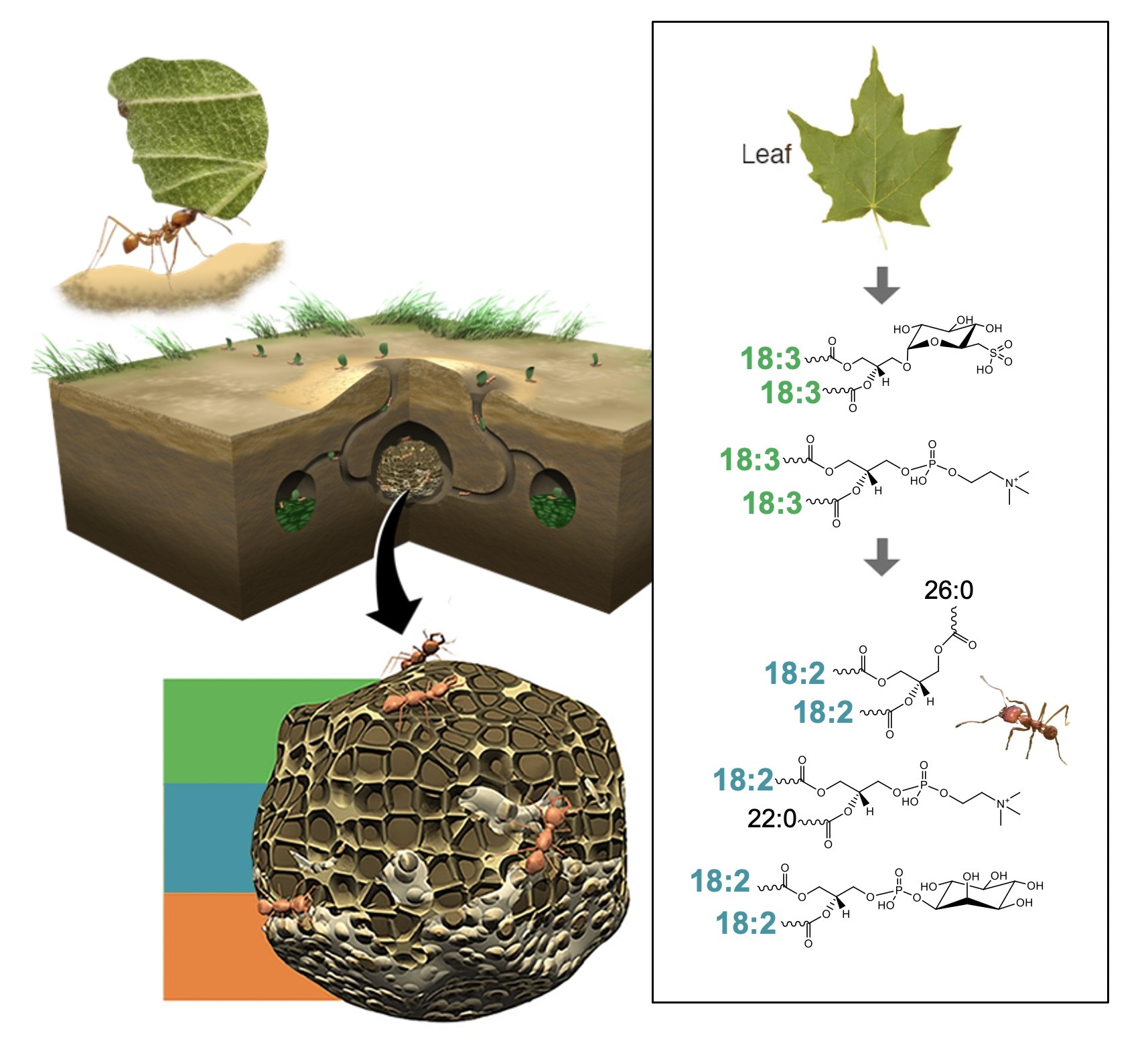
Lipids transfer energy and serve as an inter-kingdom communication tool in leaf-cutter ants’ fungal gardens.
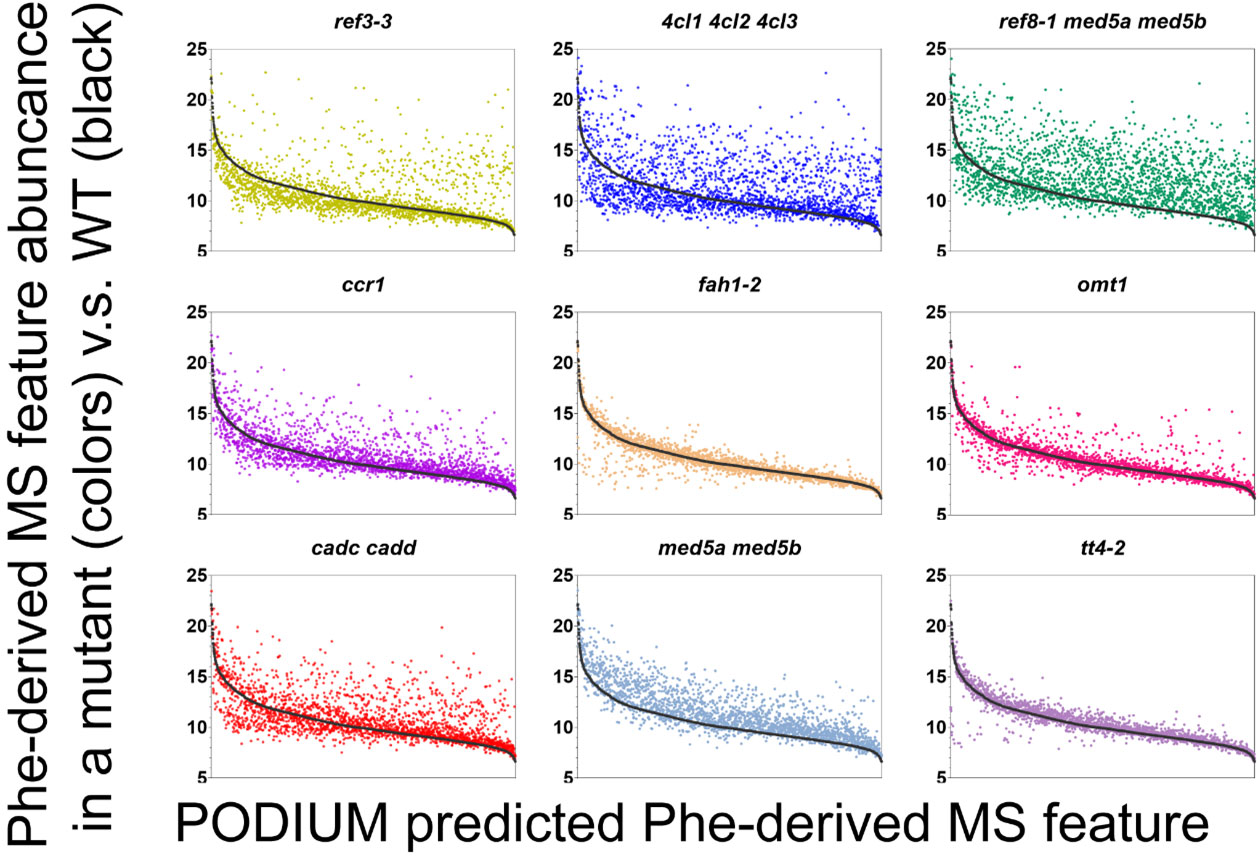
A new data pipeline identifies metabolites following heavy isotope labeling.

White-rot fungi use lignin from wood as a source of carbon.
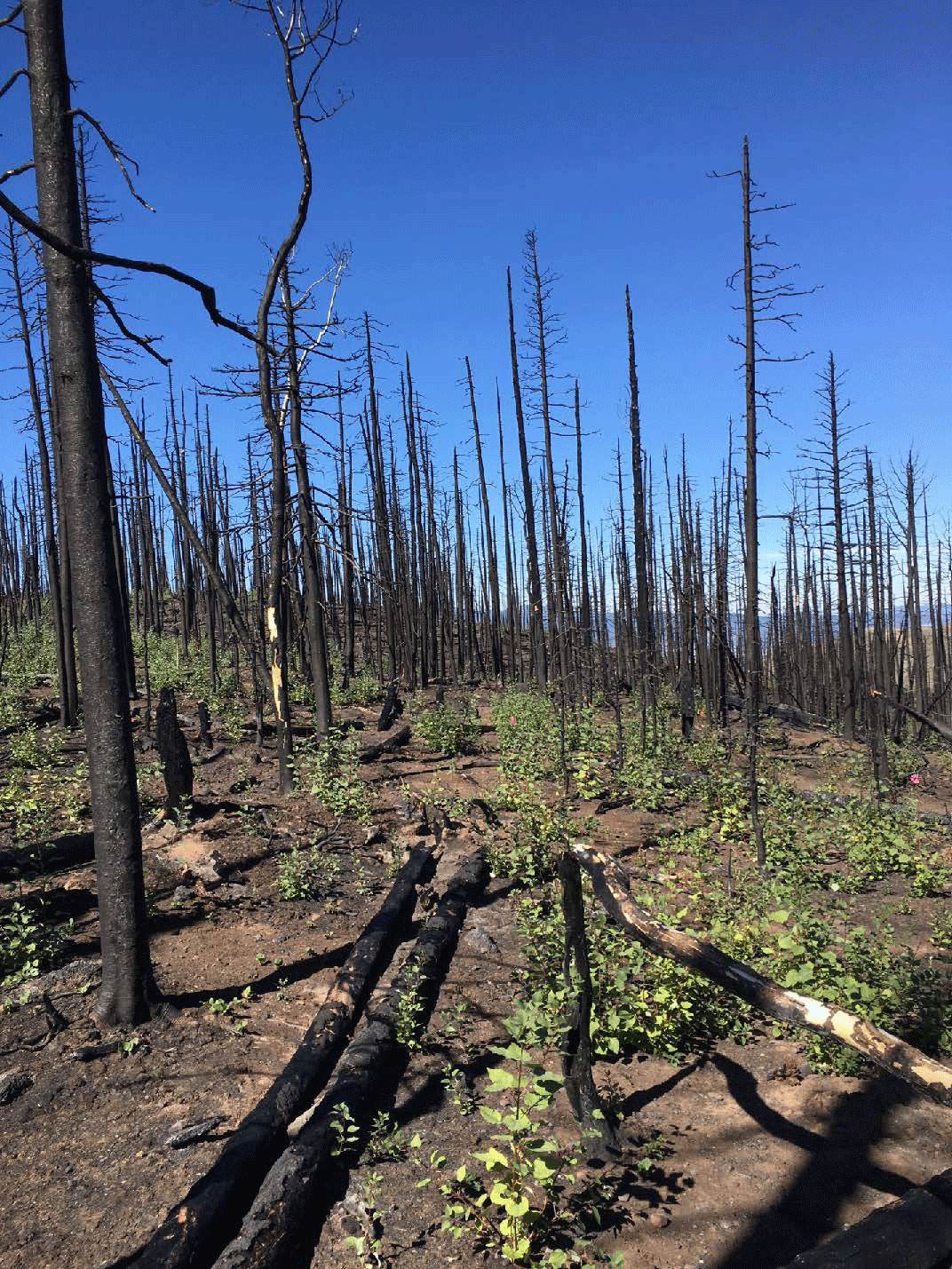

Fires increase the number of fungi when aspen groves regenerate.

Transcription of adjacent genes into a single RNA molecule is widespread in green algae, challenging understanding of gene expression in eukaryotes.